This is a package to create NONMEM Input Format (NIF) data tables from SDTM-formatted clinical study data.
The NONMEM software that is often used for population pharmacokinetic/pharmacodynamic (PK/PD) modeling expects the input data set to follow specific conventions summarized in Bauer, R.J. (2019), NONMEM Tutorial Part I: Description of Commands and Options, With Simple Examples of Population Analysis. CPT Pharmacometrics Syst. Pharmacol., 8: 525-537.
This package provides functions to sequentially aggregate drug administrations, PK/PD observations and covariates into a NONMEM-compliant analysis data set (NIF data set), and tools to explore and visualize NIF data sets.
Installation
You can install the development version of nif like this:
devtools::install_github("rstrotmann/nif", build_vignettes=TRUE)Example
Generate a NIF data set
This is a very basic example using sample SDTM data from a fictional single ascending dose study to create a NIF data set using make_nif():
library(nif)
library(tidyverse)
sdtm <- examplinib_sad
nif <- nif() %>%
add_administration(sdtm, "EXAMPLINIB", analyte = "RS2023") %>%
add_observation(sdtm, "pc", "RS2023", analyte = "RS2023")
head(nif)
#> REF ID STUDYID USUBJID AGE SEX RACE HEIGHT WEIGHT BMI
#> 1 1 1 2023000001 20230000011010001 43 0 WHITE 187.4 77 21.9256
#> 2 2 1 2023000001 20230000011010001 43 0 WHITE 187.4 77 21.9256
#> 3 3 1 2023000001 20230000011010001 43 0 WHITE 187.4 77 21.9256
#> 4 4 1 2023000001 20230000011010001 43 0 WHITE 187.4 77 21.9256
#> 5 5 1 2023000001 20230000011010001 43 0 WHITE 187.4 77 21.9256
#> 6 6 1 2023000001 20230000011010001 43 0 WHITE 187.4 77 21.9256
#> DTC TIME NTIME TAFD TAD EVID AMT ANALYTE CMT PARENT TRTDY
#> 1 2000-12-31 10:18:00 0.0 0.0 0.0 0.0 1 5 RS2023 1 RS2023 1
#> 2 2000-12-31 10:18:00 0.0 0.0 0.0 0.0 0 0 RS2023 2 RS2023 1
#> 3 2000-12-31 10:48:00 0.5 0.5 0.5 0.5 0 0 RS2023 2 RS2023 1
#> 4 2000-12-31 11:18:00 1.0 1.0 1.0 1.0 0 0 RS2023 2 RS2023 1
#> 5 2000-12-31 11:48:00 1.5 1.5 1.5 1.5 0 0 RS2023 2 RS2023 1
#> 6 2000-12-31 12:18:00 2.0 2.0 2.0 2.0 0 0 RS2023 2 RS2023 1
#> METABOLITE DOSE MDV ACTARMCD IMPUTATION DV
#> 1 FALSE 5 1 C1 NA
#> 2 FALSE 5 0 C1 0.0000
#> 3 FALSE 5 0 C1 40.7852
#> 4 FALSE 5 0 C1 48.5530
#> 5 FALSE 5 0 C1 44.0391
#> 6 FALSE 5 0 C1 34.0729In many cases, you may want to add further covariates, e.g., baseline creatinine from the LB domain:
nif <- nif %>%
mutate(COHORT = ACTARMCD) %>%
add_baseline(sdtm, "lb", "CREAT") %>%
add_bl_crcl()
#> baseline_filter for BL_CREAT set to LBBLFL == 'Y'Data exploration
The nif package provides a range of functions to explore and summarize NIF files:
summary(nif)
#> ----- NONMEM Input Format (NIF) data summary -----
#> Data from 48 subjects across one study:
#> STUDYID N
#> 2023000001 48
#>
#> Sex distribution:
#> SEX N percent
#> male 48 100
#> female 0 0
#>
#> Renal impairment class:
#> CLASS N percent
#> normal 46 95.8
#> mild 2 4.2
#> moderate 0 0
#> severe 0 0
#>
#> Treatments:
#> RS2023
#>
#> Analytes:
#> RS2023
#>
#> Subjects per dose level:
#> COHORT RS2023 N
#> C1 5 3
#> C10 500 12
#> C2 10 3
#> C3 20 3
#> C4 50 3
#> C5 100 6
#> C6 200 3
#> C7 500 6
#> C8 800 6
#> C9 1000 3
#>
#> 816 observations:
#> CMT ANALYTE N
#> 2 RS2023 816
#>
#> Sampling schedule:
#> NTIME RS2023
#> 0 X
#> 0.5 X
#> 1 X
#> 1.5 X
#> 2 X
#> 3 X
#> 4 X
#> 6 X
#> 8 X
#> 10 X
#> (7 more rows)
#>
#> Subjects with dose reductions
#> RS2023
#> 0
#>
#> Treatment duration overview:
#> PARENT min max mean median
#> RS2023 1 1 1 1
#>
#> Hash: a14674b06b234d32d526f9c20f0d0e82
#> Last DTC: 2001-03-02 11:31:00
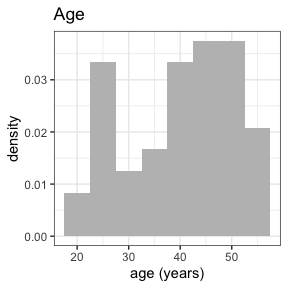
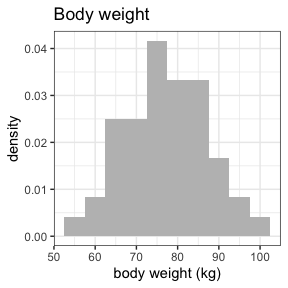
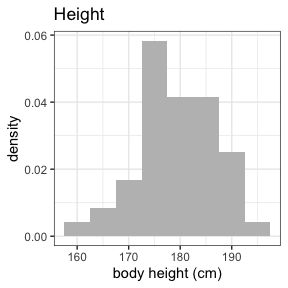
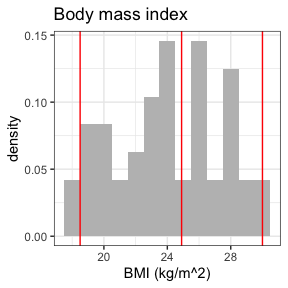
invisible(capture.output(
summary(nif) %>%
plot()
))

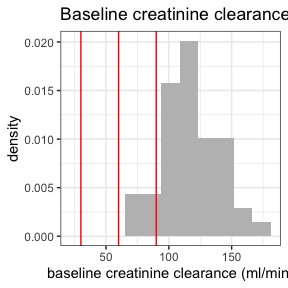
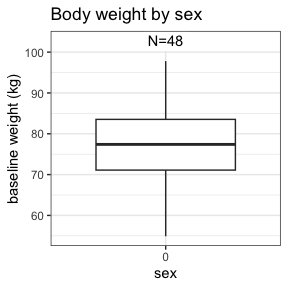
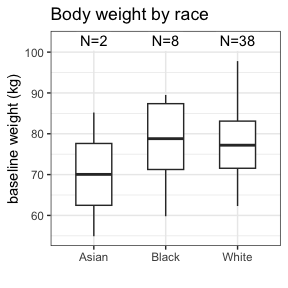
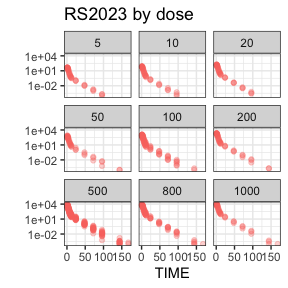
Further information
For further guidance see the help for individual functions and the project website on github pages.