INTRODUCTION
This is a basic tutorial on using the nif package to
create NONMEM Input Format (NIF) data sets from Study Data Tabulation
Model (SDTM) formatted data.
Background
Following regulatory standards, clinical study data are commonly provided in SDTM format, an observation-based data tabulation format in which logically related observations are organized into topical collections (domains). SDTM is defined and maintained by the Clinical Data Interchange Standards Consortium (CDISC).
To support typical pharmacometric analyses, data from different SDTM domains need to be aggregated into a single analysis data set. For example, demographic and pharmacokinetic concentration data from the DM and PC domains are both required to evaluate exposure by age. More complex analyses like population-level PK and PK/PD analyses may include further data, e.g., clinical laboratory, vital sign, or biomarker data.
NONMEM and other modeling software packages expect the input data provided in (long) tabular arrangement with strict requirements to the formatting and nomenclature of the variables (see, e.g., Bauer, CPT Pharmacometrics Syst. Pharmacol. (2019). The input data file for these analyses is sometimes casually referred to as a ‘NONMEM input format’ or ‘NIF’ file, hence the name of this package.
Contingent on the downstream analyses, some of the variables in the analysis data set can be easily and automatically derived from the SDTM source data, e.g., ‘DOSE’ (the administered dose) or ‘DV’ (the dependent variable for observations), or demographic covariates like ‘AGE’, ‘SEX’ or ‘RACE’. Other fields of the input data set may require study-specific considerations, for example the calculation of baseline renal or hepatic function categories, definition of specific treatment conditions by study arm, or the encoding of adverse events or concomitant medications, as categorical covariates.
While the latter variables often need manual and study-specific data
programming, the core NIF data set can in most cases be generated in a
quite standardized way. But also the manual addition of further
analysis-specific fields is substantially simplified by the functions
that the nif package provides. Often, analysis data sets
can be thus be created with only a handful of lines of code.
Even for basic NIF data sets, missing data or data inconsistencies,
specifically those relating to date and time fields, need to be resolved
to get to analysis-ready data sets. This is frequently encountered when
analyzing preliminary data from ongoing clinical studies that have not
been fully cleaned. The nif package provides a number of
standardized imputation rules to resolve these issues. More on this
point later as well in a separate vignette
(vignette("nif-imputations")).
Overall, the nif package is intended to facilitate the
creation of analysis data sets (‘NIF data sets’) from SDTM-formatted
clinical data.
Outline
The first part of this tutorial describes
how to import SDTM data into a sdtm object, and how to
explore clinical data on the SDTM level.
The second part walks through the
generation of a sample nif data set from SDTM data to
illustrate the general workflow for building analysis data sets.
Finally, the third part showcases some functions to quickly explore analysis data sets.
This tutorial contains live code that depends on the following R packages:
SDTM DATA
Importing SDTM data
In many cases, source SDTM data are provided as one file per domain, often in SAS binary data base storage format (.sas7bdat) or SAS Transport File (.xpt) format.
The SDTM data can be loaded using read_sdtm():
read_sdtm("path/to/sdtm/data")Windows users may want to provide the file path as raw string, i.e., in the form of:
read_sdtm(r"(path\to\sdtm\data)")This ensures that the backslashes in the file path are correctly captured. Note the inner parentheses around the file path!
If no domains are explicitly specified, all files with the respective extension are loaded (excluding file names that start with an underscore).
The return value of this function is a sdtm object.
SDTM objects
sdtm objects are essentially aggregates (lists) of the
SDTM domains from a particular clinical study, plus some metadata. The
easiest way of creating sdtm objects is by importing the
SDTM data using read_sdtm() as shown above.
The nif package includes sample SDTM data sets for
demonstration purposes. These data do not come from actual clinical
studies but are fully synthetic data sets from a fictional single
ascending dose (SAD) study (examplinib_sad), a fictional
food effect (FE) study (examplinib_fe), and a fictional
single-arm proof-of-concept (POC) study with multiple-dose
administrations (examplinib_poc).
Printing an sdtm object shows relevant summary
information:
examplinib_fe
#> -------- SDTM data set summary --------
#> Study 2023000400
#>
#> An open-label 2-period crossover study in healthy subjects to investigate the
#> effect of food on the pharmacokinetics of examplinib
#>
#> Data disposition
#> DOMAIN SUBJECTS OBSERVATIONS
#> dm 28 28
#> vs 28 56
#> ex 20 40
#> pc 20 1360
#> lb 28 28
#> ts 0 0
#> pp 20 360
#>
#> Arms (DM):
#> ACTARMCD ACTARM
#> AB Fasted - Fed
#> BA Fed - Fasted
#> SCRNFAIL Screen Failure
#>
#> Treatments (EX):
#> EXAMPLINIB
#>
#> PK sample specimens (PC):
#> PLASMA
#>
#> PK analytes (PC):
#> PCTEST PCTESTCD
#> RS2023 RS2023
#> RS2023487A RS2023487A
#>
#> Hash: 295fd51437d90b19b7e4d80e4c7dd51f
#> Last DTC: 2001-03-10 11:28:00High-level subject-level disposition data for a specific subject can
be extracted using subject_info():
examplinib_fe %>%
subject_info("20230004001050001")
#> [,1]
#> SITEID 105
#> SUBJID 1050001
#> ACTARM Fasted - Fed
#> ACTARMCD AB
#> RFICDTC 2000-12-26T10:05
#> RFSTDTC 2001-01-05T10:05
#> RFXSTDTC 2001-01-05T10:05
#> STUDYID 2023000400
#> USUBJID 20230004001050001
#> SEX M
#> AGE 34
#> AGEU YEARS
#> COUNTRY DEU
#> DOMAIN DM
#> ARM Fasted - Fed
#> ARMCD AB
#> RACE BLACK OR AFRICAN AMERICAN
#> ETHNIC
#> RFENDTC 2001-01-18T10:05The original SDTM data for specific domains can be retrieved from
sdtm objects with the domain() function:
domain(examplinib_sad, "dm") %>%
head(3)
#> SITEID SUBJID ACTARM ACTARMCD RFICDTC
#> 1 101 1010001 Treatment cohort 1, 5 mg examplinib C1 2000-12-21T10:18
#> 2 101 1010002 Treatment cohort 1, 5 mg examplinib C1 2000-12-21T10:30
#> 3 101 1010003 Treatment cohort 1, 5 mg examplinib C1 2000-12-21T09:22
#> RFSTDTC RFXSTDTC STUDYID USUBJID SEX AGE AGEU
#> 1 2000-12-31T10:18 2000-12-31T10:18 2023000001 20230000011010001 M 43 YEARS
#> 2 2000-12-29T10:30 2000-12-29T10:30 2023000001 20230000011010002 M 49 YEARS
#> 3 2000-12-29T09:22 2000-12-29T09:22 2023000001 20230000011010003 M 46 YEARS
#> COUNTRY DOMAIN ARM ARMCD
#> 1 DEU DM Treatment cohort 1, 5 mg examplinib C1
#> 2 DEU DM Treatment cohort 1, 5 mg examplinib C1
#> 3 DEU DM Treatment cohort 1, 5 mg examplinib C1
#> RACE ETHNIC RFENDTC
#> 1 WHITE 2000-12-31T10:18
#> 2 WHITE 2000-12-29T10:30
#> 3 BLACK OR AFRICAN AMERICAN 2000-12-29T09:22summary() generates a high-level data disposition
overview on a specific domain:
summary(domain(examplinib_sad, "pc"), silent = TRUE)
#> -------- SDTM domain summary --------
#> Study 2023000001
#> Domain PC
#> 48 subjects
#> 1632 observations
#>
#> Testcodes
#> PCTEST PCTESTCD PCSPEC
#> RS2023 RS2023 PLASMA
#> RS2023487A RS2023487A PLASMA
#>
#> Observation time points
#> PCTPT PCTPTNUM PCELTM
#> PREDOSE 0 PT0H
#> 0.5 HOURS POST-DOSE 0.5 PT0.5H
#> 1 HOURS POST-DOSE 1 PT1H
#> 1.5 HOURS POST-DOSE 1.5 PT1.5H
#> 2 HOURS POST-DOSE 2 PT2H
#> 3 HOURS POST-DOSE 3 PT3H
#> 4 HOURS POST-DOSE 4 PT4H
#> 6 HOURS POST-DOSE 6 PT6H
#> 8 HOURS POST-DOSE 8 PT8H
#> 10 HOURS POST-DOSE 10 PT10H
#> (7 more rows)
#>
#> Epochs
#> OPEN LABEL TREATMENT
#> Hash: d29adb9ffafcfd308f6b38947fefecde
#> Last DTC: 2001-03-02 12:31:00A graphical timeline overview can be generated for a specific domain by, e.g.:
plot(examplinib_fe, "dm")SDTM suggestions
SDTM data may be incomplete, e.g., when emerging data that have not yet been fully cleaned are analyzed. In addition, some study-specific data may be encoded in a non-standardized way, e.g., information on study parts, cohorts, treatment conditions, etc..
Such data fields may need study-specific considerations and manual
imputations during the creating of the analysis data set. To help
deciding which study-specific factors need to be addressed, the
nif package includes functions to explore the structure of
SDTM data.
As we will see in NIF DATA SETS, nif
objects are built sequentially from the SDTM data set. Based on the
actual data disposition, the suggest() function provide
useful code snippets for the creation of analysis data sets:
suggest(examplinib_fe)
#>
#> ── 1. Treatments ───────────────────────────────────────────────────────────────
#> There are 1 treatments (EXTRT) in EX: EXAMPLINIB. Consider adding them to the
#> nif object using `add_administration()`, see the code snippet below (replace
#> 'sdtm' with the name of your sdtm object):
#>
#> add_administration(sdtm, 'EXAMPLINIB')
#>
#> ── 2. Pharmacokinetic observations ─────────────────────────────────────────────
#> There are 2 pharmacokinetic analytes:
#>
#> PCTEST PCTESTCD
#> RS2023 RS2023
#> RS2023487A RS2023487A
#>
#> Consider adding them to the nif object using `add_observation()`, see the code
#> snippet below (replace 'sdtm' with the name of your sdtm object):
#>
#> add_observation(sdtm, 'pc', 'RS2023')
#> add_observation(sdtm, 'pc', 'RS2023487A')
#>
#> ── NTIME definition ──
#>
#> The PC domain contains multiple fields that the nominal sampling time can be
#> derived from:
#>
#> PCTPT PCTPTNUM PCELTM
#> PREDOSE 0 PT0H
#> 0.5 HOURS POST-DOSE 0.5 PT0.5H
#> 1 HOURS POST-DOSE 1 PT1H
#> 1.5 HOURS POST-DOSE 1.5 PT1.5H
#> 2 HOURS POST-DOSE 2 PT2H
#> 3 HOURS POST-DOSE 3 PT3H
#> 4 HOURS POST-DOSE 4 PT4H
#> 6 HOURS POST-DOSE 6 PT6H
#> 8 HOURS POST-DOSE 8 PT8H
#> 10 HOURS POST-DOSE 10 PT10H
#> 12 HOURS POST-DOSE 12 PT12H
#> 24 HOURS POST-DOSE 24 PT24H
#> 48 HOURS POST-DOSE 48 PT48H
#> 72 HOURS POST-DOSE 72 PT72H
#> 96 HOURS POST-DOSE 96 PT96H
#> 144 HOURS POST-DOSE 144 PT144H
#> 168 HOURS POST-DOSE 168 PT168H
#>
#> Consider specifying a suitable 'ntime_method' argument to 'add_observation()'.
#> By default, the function will attempt to extract time information from the
#> PCTPT field.
#>
#> ── 3. Study arms ───────────────────────────────────────────────────────────────
#> There are 3 study arms defined in DM:
#>
#> ACTARMCD ACTARM
#> AB Fasted - Fed
#> BA Fed - Fasted
#> SCRNFAIL Screen Failure
#>
#> Consider defining a PART or ARM variable, filtering for a particular arm, or
#> defining a covariate based on ACTARMCD.
#>
#> ── 4. Baseline covariates ──────────────────────────────────────────────────────
#> The LB domains contains creatinine (CREAT) observations. Consider adding a
#> baseline creatinine covariate, baseline creatinine clearance (BL_CRCL) and
#> baseline renal function category:
#>
#> add_baseline(sdtm, 'lb', 'CREAT')
#> add_bl_crcl()
#> add_bl_renal()In the above example, suggestions 1 and 2 provide code for the creation of administration and observation events. We will use this code directly as provided in section [Creating NIF data sets].
Suggestion 3 notes that the DM domain defines different treatment arms that should probably be included as covariates in the analysis data set. They specify the sequence of fasted and fed administrations in this study. We will deal with this in Study-specific covariates.
NIF DATA SETS
The following sections continue using the examplinib_fe
example to demonstrate how a nif object is created from the
sdtm data object.
Basic NIF file
Depending on the analysis needs, nif objects are built
in a stepwise manner, starting from an empty nif object,
sequentially adding treatment administrations, observations, and
covariate fields. The result is a data table with individual rows for
administrations and observations that follows the naming conventions
summarized in Bauer, CPT Pharmacometrics Syst. Pharmacol. (2019).
The basic nif object automatically includes standard
demographic parameters as subject-level covariates: SEX, AGE and RACE,
and baseline WEIGHT and HEIGHT are taken from the DM and VS domains,
respectively, and merged into the data set as columns of those
names:
sdtm <- examplinib_fe
nif <- nif() %>%
add_administration(sdtm, "EXAMPLINIB", analyte = "RS2023") %>%
add_observation(sdtm, "pc", "RS2023")Note that in the above, the name of the treatment, i.e., the value of
the ‘EXTRT’ field is ‘EXAMPLINIB’ while the pharmacokinetic analyte name
(‘PCTESTCD’) is ‘RS2023’. To harmonize both, the ‘analyte’ parameter in
add_administration() was set to ‘RS2023’, too.
These are the first rows of the resulting data table:
head(nif, 5)
#> REF ID STUDYID USUBJID AGE SEX RACE HEIGHT WEIGHT BMI
#> 1 1 1 2023000400 20230004001010002 53 1 WHITE 180.4 73.1 22.46179
#> 2 2 1 2023000400 20230004001010002 53 1 WHITE 180.4 73.1 22.46179
#> 3 3 1 2023000400 20230004001010002 53 1 WHITE 180.4 73.1 22.46179
#> 4 4 1 2023000400 20230004001010002 53 1 WHITE 180.4 73.1 22.46179
#> 5 5 1 2023000400 20230004001010002 53 1 WHITE 180.4 73.1 22.46179
#> DTC TIME NTIME TAFD TAD EVID AMT ANALYTE CMT PARENT TRTDY
#> 1 2001-01-05 10:05:00 0.0 0.0 0.0 0.0 1 500 RS2023 1 RS2023 1
#> 2 2001-01-05 10:05:00 0.0 0.0 0.0 0.0 0 0 RS2023 2 RS2023 1
#> 3 2001-01-05 10:35:00 0.5 0.5 0.5 0.5 0 0 RS2023 2 RS2023 1
#> 4 2001-01-05 11:05:00 1.0 1.0 1.0 1.0 0 0 RS2023 2 RS2023 1
#> 5 2001-01-05 11:35:00 1.5 1.5 1.5 1.5 0 0 RS2023 2 RS2023 1
#> METABOLITE DOSE MDV ACTARMCD IMPUTATION DV
#> 1 FALSE 500 1 AB time imputed from PCRFTDTC NA
#> 2 FALSE 500 0 AB 0.000
#> 3 FALSE 500 0 AB 4697.327
#> 4 FALSE 500 0 AB 6325.101
#> 5 FALSE 500 0 AB 6294.187Multiple analytes
To demonstrate how to add multiple analytes to a nif
object, we will temporarily switch to another built-in sample data set,
examplinib_sad. This sdtm object includes
pharmacokinetic concentration data for the M1 metabolite of ‘EXAMPLINIB’
under the PCTESTCD of ‘RS2023487A’. Note how in the below code, the two
analyte observations are added to the data set, setting the metabolite’s
name to ‘M1’, and how its relation to the parent compound is established
using the ‘parent’ parameter:
sdtm1 <- examplinib_sad
nif1 <- nif() %>%
add_administration(sdtm, "EXAMPLINIB", analyte = "RS2023") %>%
add_observation(sdtm, "pc", "RS2023") %>%
add_observation(sdtm, "pc", "RS2023487A", analyte = "M1", parent = "RS2023")In analogy to PK observations, observations from any SDTM domain,
e.g., LB, VS, MB, TR, etc., can be added in very much the same way.
Please see the documentation to add_observation() for
details. This is a powerful feature that allows effortless construction
of analysis data sets for population PK/PD modeling.
Study-specific covariates
In this simulated study, participants had received the test drug,
EXAMPLINIB, under fasted or fed conditions in a randomized sequence (see
‘ACTARM’ and ‘ACTARMCD’ in the output of suggest()), where
the ‘EPOCH’ field in the EX domain provides information on the current
treatment period. It should be noted that the way such information is
encoded in the SDTM data varies considerably. This is an example, and
the specifics of how covariate information can extracted from a SDTM
data set will differ across studies. However, nif objects
are essentially data frame objects and can thus be easily manipulated,
e.g., using functions from the dplyr package.
The following code shows how in this specific case, covariates relating to the current treatment period (‘PERIOD’) and current treatment (‘TREATMENT’) are sequentially derived and eventually used to create the ‘FASTED’ covariate.
Note that the ‘EPOCH’ field is not a standard field to be imported
from the SDTM data but can be included using the keep
parameter in add_administration():
nif <- nif() %>%
add_administration(
examplinib_fe, "EXAMPLINIB",
analyte = "RS2023", keep = "EPOCH"
) %>%
add_observation(examplinib_fe, "pc", "RS2023") %>%
mutate(PERIOD = str_sub(EPOCH, -1, -1)) %>%
mutate(TREATMENT = str_sub(ACTARMCD, PERIOD, PERIOD)) %>%
mutate(FASTED = case_when(TREATMENT == "A" ~ 1, .default = 0))These are again the first 3 lines of the resulting nif object:
head(nif, 3)
#> REF ID STUDYID USUBJID AGE SEX RACE HEIGHT WEIGHT BMI
#> 1 1 1 2023000400 20230004001010002 53 1 WHITE 180.4 73.1 22.46179
#> 2 2 1 2023000400 20230004001010002 53 1 WHITE 180.4 73.1 22.46179
#> 3 3 1 2023000400 20230004001010002 53 1 WHITE 180.4 73.1 22.46179
#> DTC TIME NTIME TAFD TAD EVID AMT ANALYTE CMT PARENT TRTDY
#> 1 2001-01-05 10:05:00 0.0 0.0 0.0 0.0 1 500 RS2023 1 RS2023 1
#> 2 2001-01-05 10:05:00 0.0 0.0 0.0 0.0 0 0 RS2023 2 RS2023 1
#> 3 2001-01-05 10:35:00 0.5 0.5 0.5 0.5 0 0 RS2023 2 RS2023 1
#> METABOLITE DOSE MDV ACTARMCD IMPUTATION DV
#> 1 FALSE 500 1 AB time imputed from PCRFTDTC NA
#> 2 FALSE 500 0 AB 0.000
#> 3 FALSE 500 0 AB 4697.327
#> EPOCH PERIOD TREATMENT FASTED
#> 1 OPEN LABEL TREATMENT 1 1 A 1
#> 2 OPEN LABEL TREATMENT 1 1 A 1
#> 3 OPEN LABEL TREATMENT 1 1 A 1DATA EXPLORATION
Data disposition
It is generally an excellent idea to explore data sets before
proceeding into more complex analyses. The nif package
provides a host of functions to this end. The following section shows
some basic examples.
The summary() function generates a general overview on
the data disposition in a nif data set:
summary(nif)
#> ----- NONMEM Input Format (NIF) data summary -----
#> Data from 20 subjects across one study:
#> STUDYID N
#> 2023000400 20
#>
#> Sex distribution:
#> SEX N percent
#> male 13 65
#> female 7 35
#>
#> Treatments:
#> RS2023
#>
#> Analytes:
#> RS2023
#>
#> Subjects per dose level:
#> RS2023 N
#> 500 20
#>
#> 680 observations:
#> CMT ANALYTE N
#> 2 RS2023 680
#>
#> Observations by NTIME:
#> NTIME RS2023
#> 0 40
#> 0.5 40
#> 1 40
#> 1.5 40
#> 2 40
#> 3 40
#> 4 40
#> 6 40
#> 8 40
#> 10 40
#> (7 more rows)
#>
#> Subjects with dose reductions
#> RS2023
#> 0
#>
#> Treatment duration overview:
#> PARENT min max mean median
#> RS2023 2 2 2 2
#>
#> Hash: 281979a9c070e4b08518ad0a83f89c75
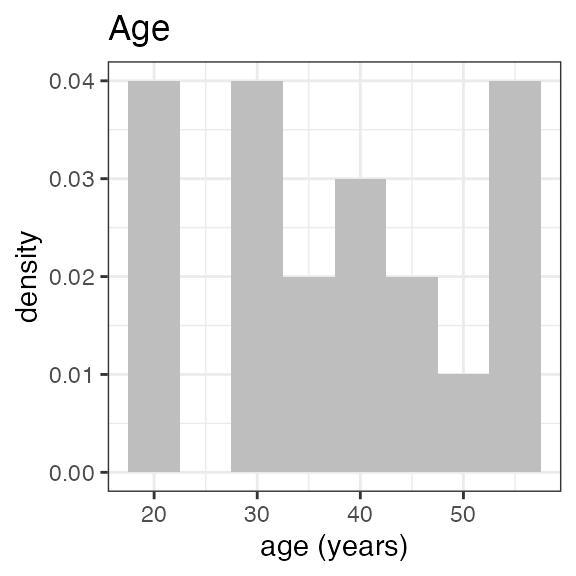
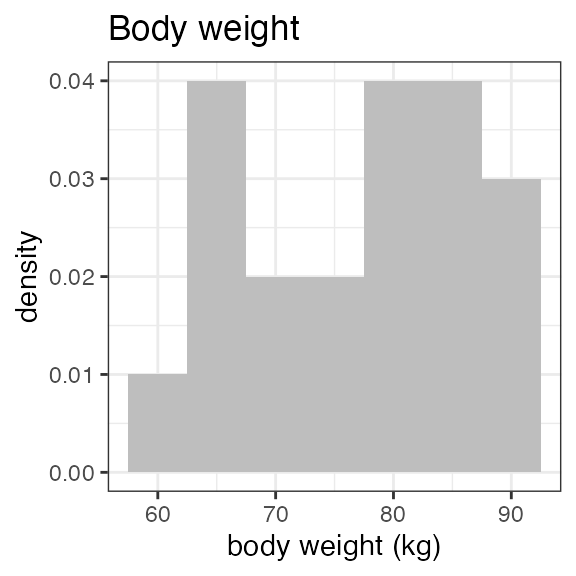
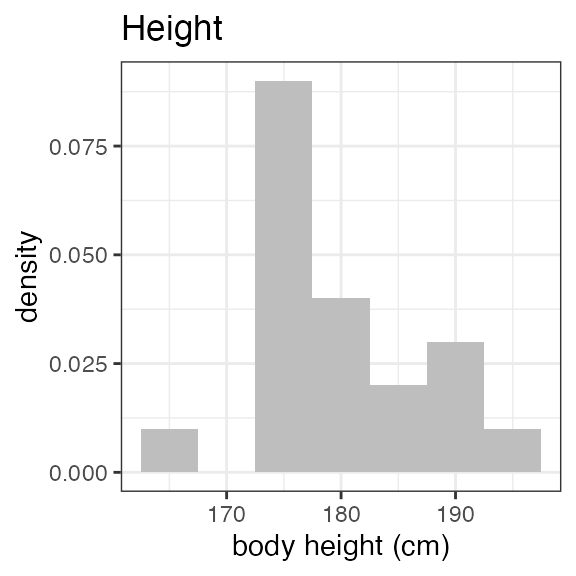
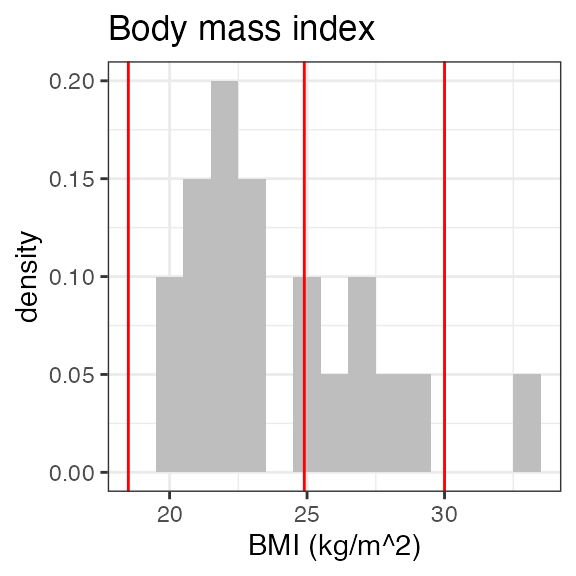
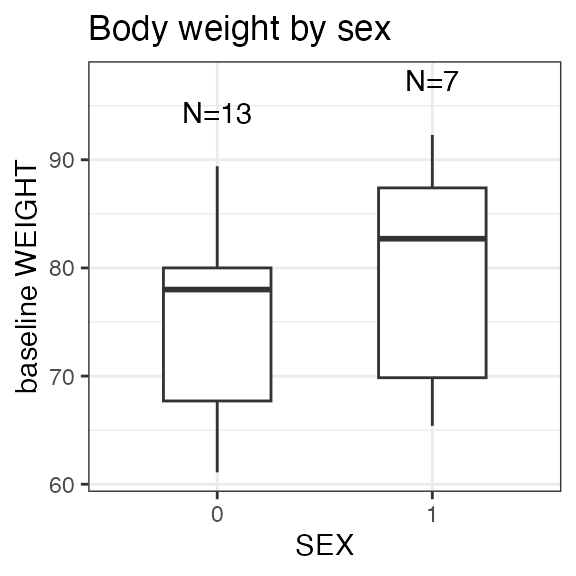
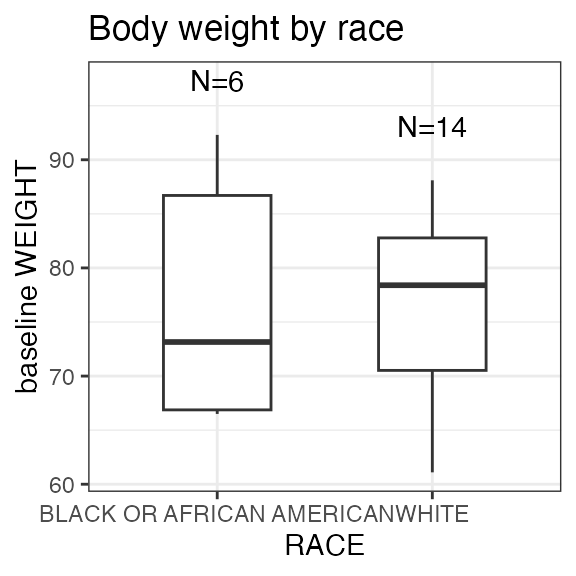
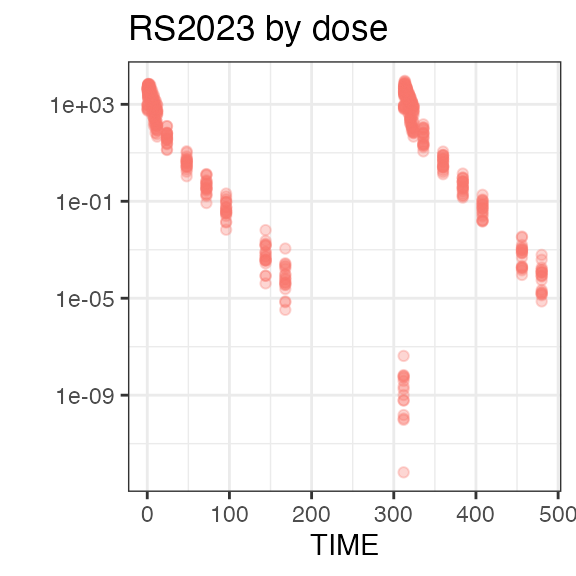
#> Last DTC: 2001-03-10 10:28:00Plotting the summary yields histograms of the baseline demographic
covariates and raw plots of the analytes over time. In the following
code, ignore the ìnvisible(capture.output()) construct
around the plot() function. Its sole purpose is to omit
some non-graphical output:
Plasma concentration data
nif objects can be easily plotted as time series charts
using the generic plot() function. While the output is a
standard ggplot2 object that can be further extended using ggplot2
functionality, the plot() function itself includes
extensive parameters to achieve the desired data visualization.
In its simplest form, plot() includes all analytes, and
uses ‘time after first dose’ (‘TAFD’) as the time metric:
plot(nif)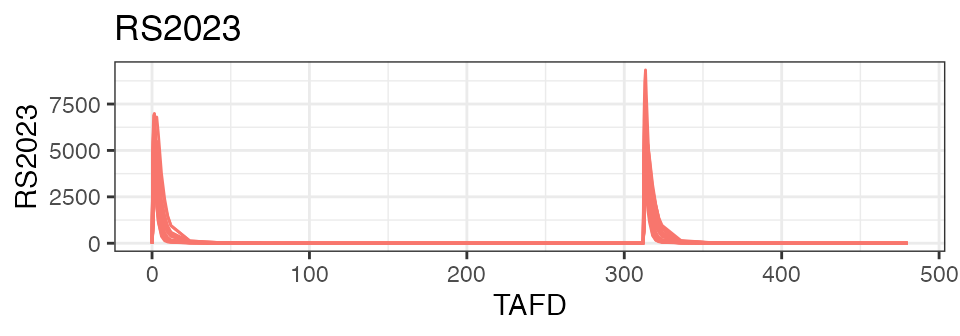
To check the integrity of the data set, it is often helpful to plot the analyte concentrations over time-after-dose (TAD):
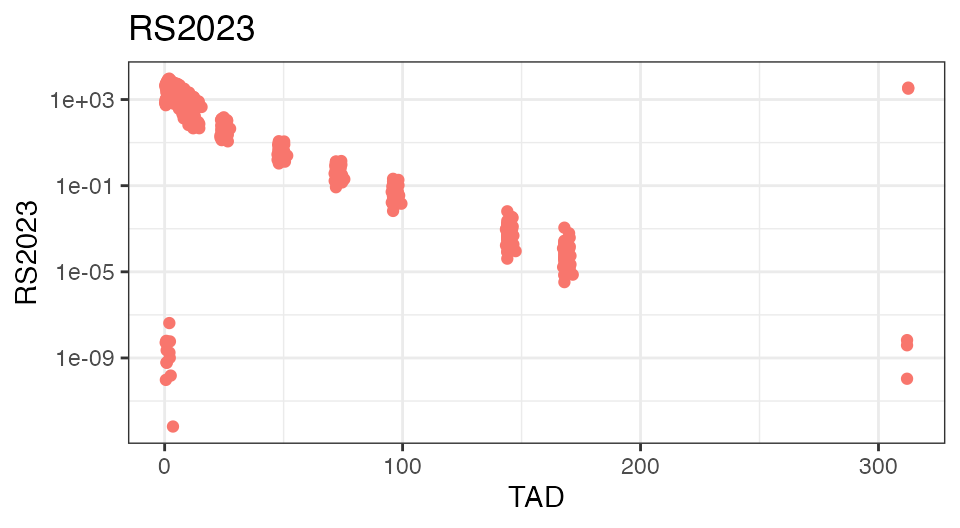
To demonstrate the food effect on Cmax and Tmax on the individual level, the below figure focuses on the first 24 hours on the linear scale and introduces coloring based on the ‘FASTED’ covariate field:
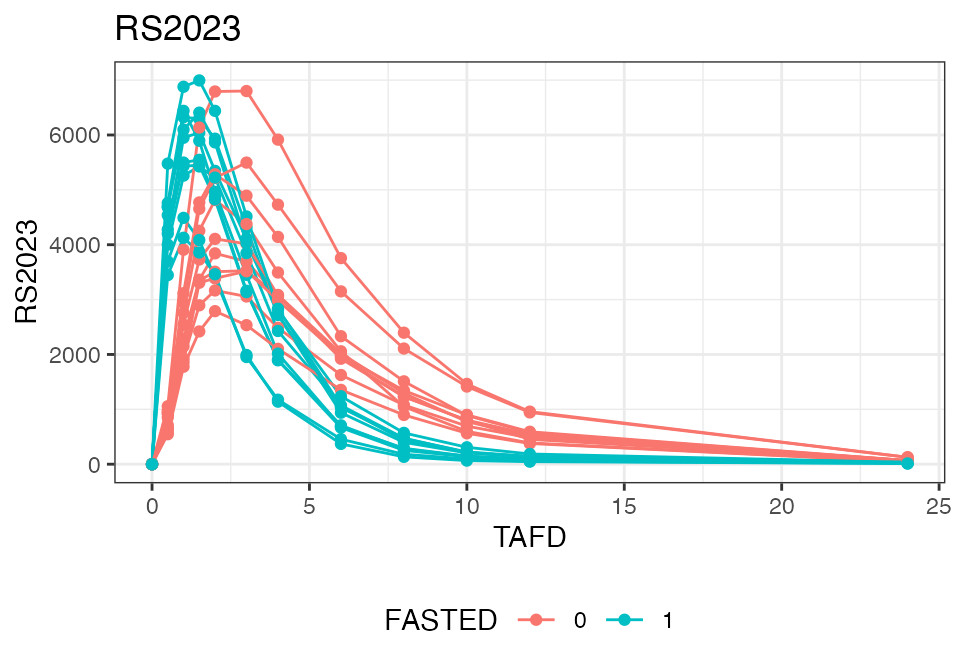
The following compares the mean plasma concentration profiles:
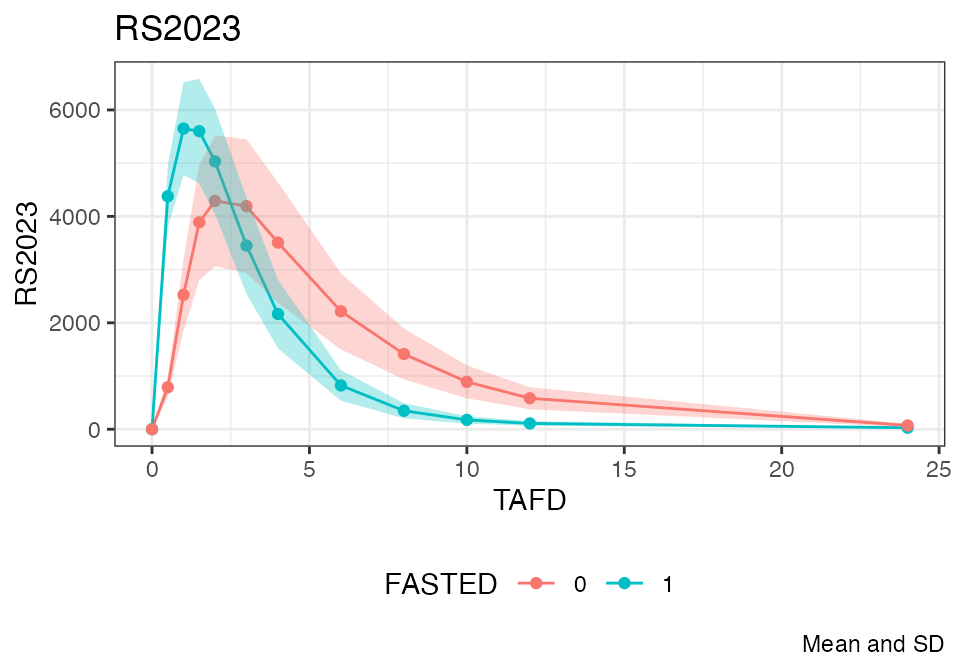
Refer to the documentation of plot.nif() for further
options.
NIF viewer
nif_viewer() is a powerful exploratory tool that lets
you interactively explore all analyte profiles on an individual level.
As the static nature of a vignette does not allow to fully appreciate
its potential, you are encouraged to test nif_viewer()
within your RStudio.
nif_viewer(nif)